 Blogs
Blogs
Dawn Dufield, PhD is the Scientific Officer for Mass Spectrometry and has been with KCAS Bio since 2018. Before joining KCAS Bio, she worked in the quantitative large and small molecule LC-MS/MS field for Pfizer for over 20 years. Dawn has been a pioneer in the Hybrid LCMS field and…
 Blogs
Blogs
Over the past decade, a continued discussion point has been the idea around analyzing samples on Ligand Binding Assay (LBA) platforms in singlet (one well) versus the standard duplicate analysis (Single sample added to two different wells). In the recent M10 Bioanalytical Method Validation Guideline issued for guidance in June…
 Blogs
Blogs
At KCAS Bio, we continuously seek ways to improve our processes with sound, logical, scientific, and business-related methods. One of these areas of evaluation has been sample preparation. We have improved various segments of bioanalysis (drying for unstable analytes, liquid handling, and plate-based assays for increased efficiency). We can also…
 Blogs
Blogs
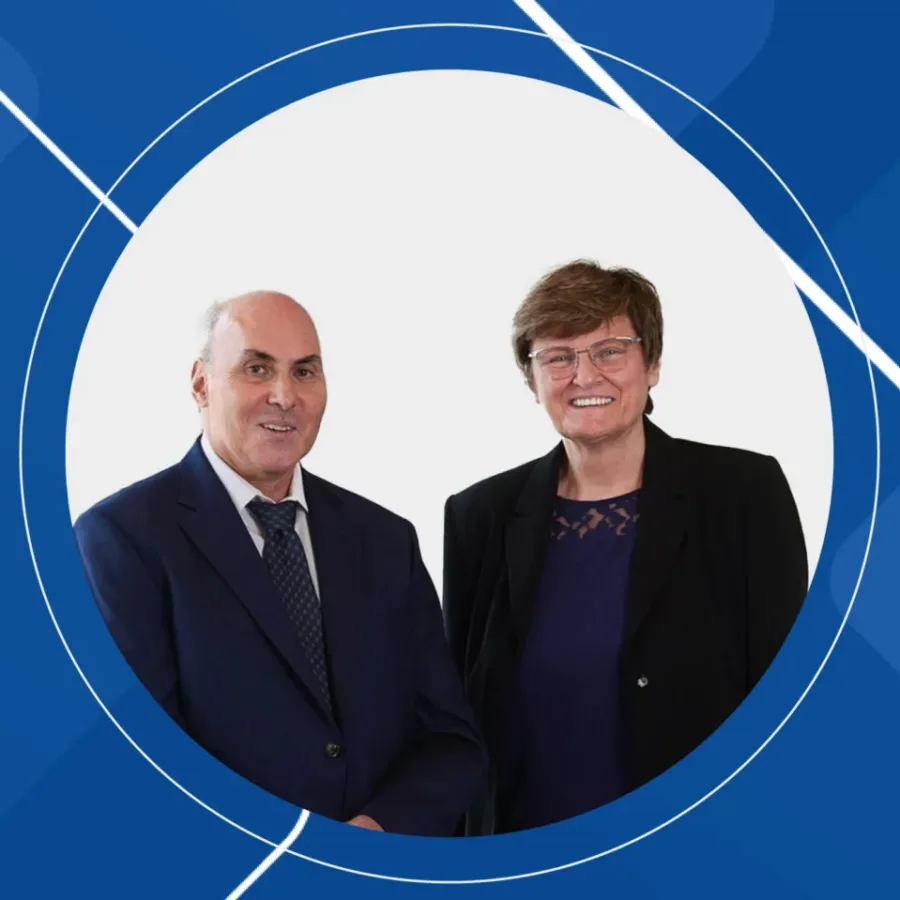
Pioneers of RNA Medicine: The Collaborative Journey of Katalin Karikó and Drew Weissman The entire world benefited from their research in 2020. Despite facing skepticism, their collaborative efforts led to groundbreaking discoveries in RNA biology and immunology. They jointly received the Nobel Prize in Physiology or Medicine in 2023.
 Blogs
Blogs
KCAS Bio SAS (Active Biomarkers SAS) based in Lyon France, a full-service bioanalytical and biomarkers solutions provider is very pleased and proud to share that it successfully opened a new Good Laboratory Practice (GLP) Test Facility to provide GLP-grade bioanalytical methods in the context of preclinical studies. Since November 2023,…
 Blogs
Blogs
As an emerging and growing application, spectral flow offers capabilities beyond what is possible with conventional flow. Its unique capabilities make it a powerful tool for researchers aiming to extract more information from each sample and push the boundaries of discovery. Join us in this post as we explore…
 Blogs
Blogs
In the scientific field, the often neglected fifth sense, the sense of smell, contributes to broadening our understanding of various phenomena, particularly in the field of biomarkers.Here, we will guide you along our lines of thought on the importance of our olfactory sense, and how it contributes to scientific exploration…
 Blogs
Blogs
For the last decade or so, the outsourcing of crucial research functions by pharmaceutical and biotechnology companies has become increasingly common and has resulted in the rapid growth of the Contract Research Organization (CRO) industry. But the selection of the best CRO partner to work on your critical research programs…
 Blogs
Blogs
Rosalind Franklin: Unveiling the DNA Helix and Beyond She is known for the celebrated photo 51 She is famous for her X-rays An American university and an English research institute bear her name Her name is Rosalind Franklin. Her crucially important X-ray crystallography work was key to…
 Blogs
Blogs
Flow Cytometry has been an invaluable tool for scientific advancement in immunology, oncology, stem cell research, infectious disease, vaccines, and drug development research. As a versatile and powerful technique, it can be used in various ways and tailored to meet your research needs. Applications include precise evaluation of cell…
 Blogs
Blogs
You might be asking what type of laboratory my biomarker assay requires or what level of qualification or validation my assay needs. These are complex questions which there continues to be a significant amount of misunderstanding around. Fortunately, KCAS Bio has extensive experience with both biomarker development and qualification/validation to…
 Blogs
Blogs
Immunoaffinity LC-MS/MS or hybrid LC-MS/MS is an approach that has been gaining a lot of traction in the past several years. The technique allows the quantitation of low-level analytes (proteins or peptides in many cases) in biological matrices with exquisite selectivity and sensitivity. Many pharmaceutical companies are trying to expand…